Representation learning to advance multi-institutional studies with electronic health record data from US and France
Representation learning to advance multi-institutional studies with electronic health record data from US and France
References
- Liao, K. P. et al. Development of phenotype algorithms using electronic medical records and incorporating natural languageprocessing. BMJ 350, h1885 https://doi.org/10.1136/bmj.h1885 (2015).
- Wang, L. et al. Stratification of Alzheimer’s disease patients using knowledge-guided unsupervised latent factor clustering with electronic health record data. Preprint at Dec 26 https://doi.org/10.1101/2024.12.23.24319588 (2024).
- Doshi-Velez, F., Ge, Y. & Kohane, I. Comorbidity clusters in autism spectrum disorders: an electronic health record time-series analysis. Pediatrics 133, e54–e63 (2014).
Google Scholar
- Sheu, Y. -h. et al. An efficient landmark model for prediction of suicide attempts in multiple clinical settings. Psychiatry Res. 323, 115175 (2023).
Google Scholar
- Federico, P. et al. Gnaeus: Utilizing clinical guidelines for knowledge-assisted visualisation of EHR cohorts. In Roberts, J. C. & Bertini, E. (eds.) 6th International EuroVis Workshop on Visual Analytics, EuroVA@EuroVis 2015, Cagliari, Sardinia, Italy, May 25-26, 2015, 79–83 (Eurographics Association, 2015).
- Ferté, T., Jouhet, V., Griffier, R., Hejblum, B. P. & Thiébaut, R. The benefit of augmenting open data with clinical data-warehouse EHR for forecasting SARS-CoV-2 hospitalizations in Bordeaux area, France. JAMIA Open 5, ooac086 (2022).
Google Scholar
- Wen, J. et al. Multimodal representation learning for predicting molecule–disease relations. Bioinformatics 39, btad085 (2023).
Google Scholar
- Cai, T., Xia, D., Zhang, L. & Zhou, D. Consensus knowledge graph learning via multi-view sparse low rank block model. Preprint at https://doi.org/10.48550/arXiv.2209.13762 (2022).
- Hur, K. et al. Unifying heterogeneous electronic health records systems via text-based code embedding. In Proc. Conference on Health, Inference, and Learning, Vol. 174 of Proc. of Machine Learning Research, (eds. Flores, G., Chen, G. H., Pollard, T., Ho, J. C. & Naumann, T.) 183–203 (PMLR, 2022).
- Molaei, S. et al. Federated learning for heterogeneous electronic health records utilising augmented temporal graph attention networks. In Proc. International Conference on Artificial Intelligence and Statistics, 1342–1350 (PMLR, 2024).
- Thakur, A. et al. Knowledge abstraction and filtering based federated learning over heterogeneous data views in healthcare. NPJ Digit. Med. 7, 283 (2024).
Google Scholar
- Centre for Disease Control and Prevention et al. International classification of diseases, ninth revision (ICD-9). Cincinnati, Ohio: National Center for Health Statistics (1979).
- McDonald, C. J. et al. Loinc, a universal standard for identifying laboratory observations: a 5-year update. Clin. Chem. 49, 624–633 (2003).
Google Scholar
- Chen, M. et al. Privacy protection and intrusion avoidance for cloudlet-based medical data sharing. IEEE Trans. Cloud Comput. 8, 1274–1283 (2016).
Google Scholar
- Sheller, M. et al. Federated learning in medicine: facilitating multi-institutional collaborations without sharing patient data. Sci. Rep. 10, (2020).
- Mikolov, T., Sutskever, I., Chen, K., Corrado, G. S. & Dean, J. Distributed representations of words and phrases and their compositionality. Adv. Neural Inf. Process. Syst. 26, (2013).
- Pennington, J., Socher, R. & Manning, C. D. Glove: global vectors for word representation. In Proc. 2014 Conference on Empirical Methods in Natural Language Processing, 1532–1543 (ACL, 2014).
- Wang, Z., Zhang, J., Feng, J. & Chen, Z. Knowledge graph embedding by translating on hyperplanes. In Proc. AAAI Conference on Artificial Intelligence, Vol. 28 (AAAI, 2014).
- Balažević, I., Allen, C. & Hospedales, T. Tucker: tensor factorization for knowledge graph completion. In Proc. 2019 Conference on Empirical Methods in Natural Language Processing and the 9th International Joint Conference on Natural Language Processing (EMNLP-IJCNLP), 5185–5194 (ACL, 2019).
- Yuan, Z. et al. CODER: knowledge-infused cross-lingual medical term embedding for term normalization. J. Biomed. Inform. 126, 103983 (2022).
Google Scholar
- Lin, Y., Lu, K., Yu, S., Cai, T. & Zitnik, M. Multimodal learning on graphs for disease relation extraction. J. Biomed. Inform. 143, 104415 (2023).
Google Scholar
- Bodenreider, O. The Unified Medical Language System (UMLS): integrating biomedical terminology. Nucleic Acids Res. 32, D267–D270 (2004).
Google Scholar
- Liu, F., Shareghi, E., Meng, Z., Basaldella, M. & Collier, N. Self-alignment pretraining for biomedical entity representations. In Proc. 2021 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies, 4228–4238 (ACL, 2021).
- Maldonado, R., Yetisgen, M. & Harabagiu, S. M. Adversarial learning of knowledge embeddings for the Unified Medical Language System. AMIA Summits Transl. Sci. Proc. 2019, 543 (2019).
Google Scholar
- Michalopoulos, G., Wang, Y., Kaka, H., Chen, H. & Wong, A. UmlsBERT: clinical domain knowledge augmentation of contextual embeddings using the Unified Medical Language System Metathesaurus. In Proc. 2021 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies, 1744–1753 (ACL, 2021).
- Piya, F. L., Gupta, M. & Beheshti, R. HealthGAT: node classifications in electronic health records using graph attention networks. In Proc. 2024 IEEE/ACM Conference on Connected Health: Applications, Systems and Engineering Technologies (CHASE), 132–141 (IEEE, 2024).
- Choi, E. et al. Multi-layer representation learning for medical concepts. In Proc. 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, 1495–1504 (ACM, 2016).
- Kartchner, D., Christensen, T., Humpherys, J. & Wade, S. Code2vec: embedding and clustering medical diagnosis data. In Proc. 2017 IEEE International Conference on Healthcare Informatics, 386–390 (IEEE, 2017).
- Hong, C. et al. Clinical knowledge extraction via sparse embedding regression (KESER) with multi-center large scale electronic health record data. NPJ Digit. Med. 4, 151 (2021).
Google Scholar
- Zhou, D. et al. Multiview incomplete knowledge graph integration with application to cross-institutional EHR data harmonization. J. Biomed. Inform. 133, 104147 (2022).
Google Scholar
- Gan, Z. et al. ARCH: large-scale knowledge graph via aggregated narrative codified health records analysis. J. Biomed. Inform. 162, 104761 (2025).
- Wang, K., Chen, N. & Chen, T. Joint medical ontology representation learning for healthcare predictions. In Proc. 2020 International Joint Conference on Neural Networks (IJCNN), 1–7 (IEEE, 2020).
- Ying, H., Zhao, Z., Zhao, Y., Zeng, S. & Yu, S. CoRTEx: contrastive learning for representing terms via explanations with applications on constructing biomedical knowledge graphs. J. Am. Med. Inform. Assoc. 31, 1912–1920 (2024).
Google Scholar
- Gao, Y. et al. Leveraging medical knowledge graphs into large language models for diagnosis prediction: design and application study. JMIR AI 4, e58670 (2025).
Google Scholar
- Cai, T., Huang, F., Nakada, R., Zhang, L. & Zhou, D. Contrastive learning on multimodal analysis of electronic health records. Preprint at https://doi.org/10.48550/arXiv.2403.14926 (2024).
- Levy, O. & Goldberg, Y. Neural word embedding as implicit matrix factorization. Adv. Neural Inf. Process. Syst. 27, (2014).
- Gu, Y. et al. Domain-specific language model pretraining for biomedical natural language processing. ACM Trans. Comput. Healthc. 3, 1–23 (2021).
Google Scholar
- Lee, J. et al. Biobert: a pre-trained biomedical language representation model for biomedical text mining. Bioinformatics 36, 1234–1240 (2020).
Google Scholar
- Chen, J. et al. M3-embedding: Multi-linguality, multi-functionality, multi-granularity text embeddings through self-knowledge distillation. Findings of the Association for Computational Linguistics: ACL 2024, 2318–2335 (Association for Computational Linguistics, Bangkok, Thailand, 2024).
- Cipriani, A. et al. Comparative efficacy and acceptability of antimanic drugs in acute mania: a multiple-treatments meta-analysis. Lancet 378, 1306–1315 (2011).
Google Scholar
- Arvanitis, L. A. & Miller, B. G. Multiple fixed doses of “Seroquel” (Quetiapine) in patients with acute exacerbation of schizophrenia: a comparison with Haloperidol and placebo. Biol. Psychiatry 42, 233–246 (1997).
Google Scholar
- Ismail, Z. et al. Psychosis in Alzheimer disease-mechanisms, genetics and therapeutic opportunities. Nat. Rev. Neurol. 18, 131–144 (2022).
Google Scholar
- Liu, J., Chang, L., Song, Y., Li, H. & Wu, Y. The role of NMDA receptors in Alzheimer’s disease. Front. Neurosci. 13, 43 (2019).
Google Scholar
- Tariot, P. N. et al. Memantine treatment in patients with moderate to severe Alzheimer disease already receiving donepezil: a randomized controlled trial. J. Am. Med. Inform. Assoc. 291, 317–324 (2004).
Google Scholar
- Anthropic. Introducing the next generation of claude https://www.anthropic.com/news/claude-3-family (2024).
- Meta AI. The LLaMA 4 herd: The beginning of a new era of natively multimodal AI innovation (2025) https://ai.meta.com/blog/llama-4-multimodal-intelligence/ Accessed: 2025-Apr-07.
- OMOP. Standardized data: The OMOP common data model (2021) https://www.ohdsi.org/data-standardization/. Accessed: Jun, 2025.
- Wen, J. et al. DOME: directional medical embedding vectors from electronic health records. J. Biomed. Inform. 162, 104768 (2025).
Google Scholar
- Chen, L. et al. Graph optimal transport for cross-domain alignment. In Proc. International Conference on Machine Learning, 1542–1553 (PMLR, 2020).
- Veličković, P. et al. Graph attention networks. In Proc. International Conference on Learning Representations (ICLR, 2018).
- Gori, M., Monfardini, G. & Scarselli, F. A new model for learning in graph domains. In Proc. 2005 IEEE International Joint Conference on Neural Networks, Vol. 2, 729–734 (IEEE, 2005).
- Johnson, A. et al. MIMIC-IV (version 0.4). PhysioNet. (2020) https://physionet.org/content/mimiciv/0.4/. Accessed: June, 2025.
- Bousquet, C., Trombert, B., Souvignet, J., Sadou, E. & Rodrigues, J.-M. Evaluation of the CCAM hierarchy and semi structured code for retrieving relevant procedures in a hospital case mix database. In Proc.AMIA Annual Symposium Proceedings, Vol. 2010, 61 (AMIA, 2010).
- Beam, A. L. et al. Clinical concept embeddings learned from massive sources of multimodal medical data. In Proc. Pacific Symposium on Biocomputing, Vol. 25, 295–306 (PSB, 2020).
- Shin, H.-C. et al. BioMegatron: larger biomedical domain language model. In Proc. 2020 Conference on Empirical Methods in Natural Language Processing (EMNLP), 4700–4706 (ACL, 2020).
- Wang, X., Han, X., Huang, W., Dong, D. & Scott, M. R. Multi-similarity loss with general pair weighting for deep metric learning. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition, 5022–5030 (IEEE Computer Society, 2019).
- Li, L. et al. Identification of type 2 diabetes subgroups through topological analysis of patient similarity. Sci. Transl. Med. 7, 311ra174–311ra174 (2015).
Google Scholar
- Landi, I. et al. Deep representation learning of electronic health records to unlock patient stratification at scale. NPJ Digit. Med. 3, 96 (2020).
Google Scholar
- Van der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, (2008).
- Garst, S. & Reinders, M. Federated k-means clustering. In Proc.International Conference on Pattern Recognition, 107–122 (Springer, 2024).
- Armstrong, M. J., Song, S., Kurasz, A. M. & Li, Z. Predictors of mortality in individuals with dementia in the National Alzheimer’s Coordinating Center. J. Alzheimer’s. Dis. 86, 1935–1946 (2022).
Google Scholar
- Zheng, X., Wang, S., Huang, J., Li, C. & Shang, H. Predictors for survival in patients with Alzheimer’s disease: a large comprehensive meta-analysis. Transl. Psychiatry 14, 184 (2024).
Google Scholar
- Abdelnour, C. et al. Perspectives and challenges in patient stratification in Alzheimer’s disease. Alzheimer’s. Res. Ther. 14, 112 (2022).
Google Scholar
- Han, E., Kharrazi, H., Shi, L. et al. Identifying predictors of nursing home admission by using electronic health records and administrative data: scoping review. JMIR Aging 6, e42437 (2023).
Google Scholar
- Favril, L., Yu, R., Uyar, A., Sharpe, M. & Fazel, S. Risk factors for suicide in adults: systematic review and meta-analysis of psychological autopsy studies. BMJ Ment. Health 25, 148–155 (2022).
Google Scholar
- Sutar, R., Kumar, A. & Yadav, V. Suicide and prevalence of mental disorders: a systematic review and meta-analysis of world data on case-control psychological autopsy studies. Psychiatry Res. 329, 115492 (2023).
- Fazel, S. & Runeson, B. Suicide. N. Engl. J. Med. 382, 266–274 (2020).
Google Scholar
- Lee, D., Jiang, X. & Yu, H. Harmonized representation learning on dynamic EHR graphs. J. Biomed. Inform. 106, 103426 (2020).
Google Scholar
- Panickan, V. A., CELEHS & Tong, H. celehs/game: representation learning to advance multi-institutional studies with electronic health record data https://github.com/celehs/GAME (2026).
Download references
Sign in to highlight and annotate this article

Conversation starters
Daily AI Digest
Get the top 5 AI stories delivered to your inbox every morning.
More about
france
Docling Studio — open-source visual inspection tool for Docling pipelines
Hey everyone I built Docling Studio , an open-source visual inspection layer for Docling. The problem: if you’ve used Docling, you know the extraction engine is powerful — but validating outputs means digging through JSON and mentally mapping bounding box coordinates back to the original pages. No visual feedback loop. What Docling Studio does: Upload a PDF, configure your pipeline (OCR engine, table extraction, enrichment) Run the conversion Visually inspect every detected element — bounding boxes overlaid on original pages, element types, content preview on click Two modes: local (embedded Docling) or remote (Docling Serve) Stack: Vue 3 / TypeScript + FastAPI / Python, fully Dockerized (multi-arch), 180+ tests. Why it matters for RAG workflows: without seeing what Docling extracts, it’s
Knowledge Map
Connected Articles — Knowledge Graph
This article is connected to other articles through shared AI topics and tags.
More in Countries
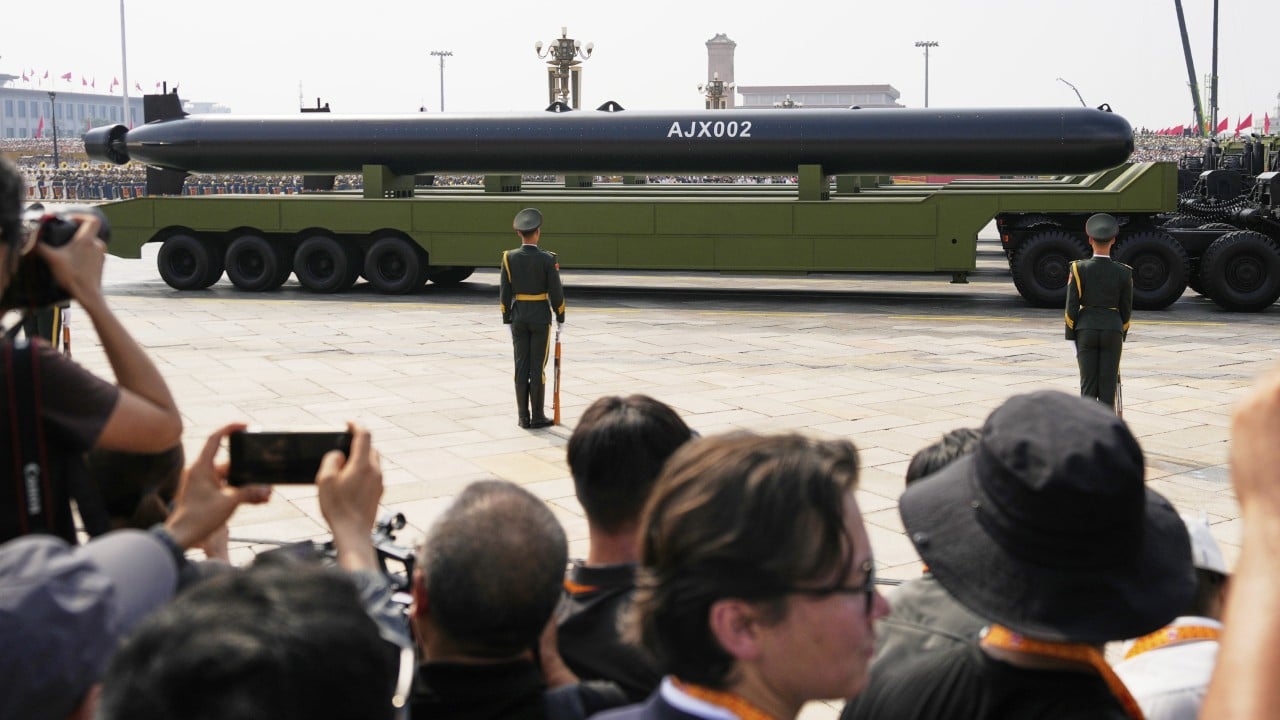

China not targeting US West Coast with ultra-large underwater drones: lead scientist
China’s unmanned submersibles now rank as the world’s largest, with last year’s military parade showcasing two models (HSU001 and AJX002) approaching 20 metres (66 feet) in length. Satellite imagery analysed by Western media also revealed a classified variant exceeding 40 metres at a naval installation, triggering international concern – particularly in the United States. These dimensions created a brand new class of drones known as extra-extra-large uncrewed underwater vehicles (XXLUUVs). They...






Discussion
Sign in to join the discussion
No comments yet — be the first to share your thoughts!