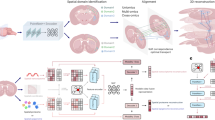
Network model for alignment, stitching and slice-to-volume 3D reconstruction of large-scale spatially resolved slices
Network model for alignment, stitching and slice-to-volume 3D reconstruction of large-scale spatially resolved slices
References
- Marx, V. Method of the year: spatially resolved transcriptomics. Nat. Methods 18, 9–14 (2021).
Google Scholar
- Ståhl, P. L. et al. Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science 353, 78–82 (2016).
Google Scholar
- Vickovic, S. et al. High-definition spatial transcriptomics for in situ tissue profiling. Nat. Methods 16, 987–990 (2019).
Google Scholar
- Marco Salas, S. et al. Optimizing xenium in situ data utility by quality assessment and best-practice analysis workflows. Nat. Methods 22, 813–823 (2025).
- Stickels, R. R. et al. Highly sensitive spatial transcriptomics at near-cellular resolution with slide-seqv2. Nat. Biotechnol. 39, 313–319 (2021).
Google Scholar
- Wei, X. et al. Single-cell stereo-seq reveals induced progenitor cells involved in axolotl brain regeneration. Science 377, eabp9444 (2022).
Google Scholar
- Andersson, A. et al. Spatial deconvolution of HER2-positive breast cancer delineates tumor-associated cell type interactions. Nat. Commun. 12, 6012 (2021).
Google Scholar
- Berglund, E. et al. Spatial maps of prostate cancer transcriptomes reveal an unexplored landscape of heterogeneity. Nat. Commun. 9, 2419 (2018).
Google Scholar
- Moncada, R. et al. Integrating microarray-based spatial transcriptomics and single-cell RNA-seq reveals tissue architecture in pancreatic ductal adenocarcinomas. Nat. Biotechnol. 38, 333–342 (2020).
Google Scholar
- Maynard, K. R. et al. Transcriptome-scale spatial gene expression in the human dorsolateral prefrontal cortex. Nat. Neurosci. 24, 425–436 (2021).
Google Scholar
- Farah, E. N. et al. Spatially organized cellular communities form the developing human heart. Nature 627, 854–864 (2024).
Google Scholar
- Maniatis, S. et al. Spatiotemporal dynamics of molecular pathology in amyotrophic lateral sclerosis. Science 364, 89–93 (2019).
Google Scholar
- Cable, D. M. et al. Cell type-specific inference of differential expression in spatial transcriptomics. Nat. Methods 19, 1076–1087 (2022).
Google Scholar
- Wu, L. et al. An invasive zone in human liver cancer identified by stereo-seq promotes hepatocyte–tumor cell crosstalk, local immunosuppression and tumor progression. Cell Res. 33, 585–603 (2023).
- Hu, J. et al. Spagcn: Integrating gene expression, spatial location and histology to identify spatial domains and spatially variable genes by graph convolutional network. Nat. Methods 18, 1342–1351 (2021).
Google Scholar
- Zhao, E. et al. Spatial transcriptomics at subspot resolution with bayesspace. Nat. Biotechnol. 39, 1375–1384 (2021).
Google Scholar
- Cable, D. M. et al. Robust decomposition of cell type mixtures in spatial transcriptomics. Nat. Biotechnol. 40, 517–526 (2022).
Google Scholar
- Li, J., Chen, S., Pan, X., Yuan, Y. & Shen, H.-B. Cell clustering for spatial transcriptomics data with graph neural networks. Nat. Comput Sci. 2, 399–408 (2022).
Google Scholar
- Dong, K. & Zhang, S. Deciphering spatial domains from spatially resolved transcriptomics with an adaptive graph attention auto-encoder. Nat. Commun. 13, 1–12 (2022).
Google Scholar
- Wang, Y., Liu, Z. & Ma, X. Mucst: restoring and integrating heterogeneous morphology images and spatial transcriptomics data with contrastive learning. Genome Med 17, 21 (2025).
Google Scholar
- Ueda, H. R. et al. Tissue clearing and its applications in neuroscience. Nat. Rev. Neurosci. 21, 61–79 (2020).
Google Scholar
- Richardson, D. S. & Lichtman, J. W. Clarifying tissue clearing. Cell 162, 246–257 (2015).
Google Scholar
- Xie, P. et al. Digital reconstruction of full embryos during early mouse organogenesis. Cell 188, 4754–4772 (2025).
- Zeira, R., Land, M., Strzalkowski, A. & Raphael, B. J. Alignment and integration of spatial transcriptomics data. Nat. Methods 19, 567–575 (2022).
Google Scholar
- Liu, X., Zeira, R. & Raphael, B. J. Partial alignment of multislice spatially resolved transcriptomics data. Genome Res 33, 1124–1132 (2023).
Google Scholar
- Klein, D. et al. Mapping cells through time and space with Moscot. Nature 638, 1065–1075 (2025).
- Xia, C.-R., Cao, Z.-J., Tu, X.-M. & Gao, G. Spatial-linked alignment tool (slat) for aligning heterogenous slices. Nat. Commun. 14, 7236 (2023).
Google Scholar
- Gao, J., Huang, X. & Li, J. Unsupervised graph alignment with Wasserstein distance discriminator. In Proceedings of the 27th ACM SIGKDD Conference on Knowledge Discovery & Data Mining, 426–435 (2021).
- Li, H. et al. Santo: a coarse-to-fine alignment and stitching method for spatial omics. Nat. Commun. 15, 6048 (2024).
Google Scholar
- Xu, H. et al. Spacel: deep learning-based characterization of spatial transcriptome architectures. Nat. Commun. 14, 7603 (2023).
Google Scholar
- Clifton, K. et al. Stalign: Alignment of spatial transcriptomics data using diffeomorphic metric mapping. Nat. Commun. 14, 8123 (2023).
Google Scholar
- Tang, Z. et al. Search and match across spatial omics samples at single-cell resolution. Nat. Methods 21, 1818–1829 (2024).
- Qiu, X. et al. Spatiotemporal modeling of molecular holograms. Cell 187, 7351–7373 (2024).
Google Scholar
- Tajdari, F., Huysmans, T., Yang, Y. & Song, Y. Feature preserving non-rigid iterative weighted closest point and semi-curvature registration. IEEE Trans. Image Process. 31, 1841–1856 (2022).
Google Scholar
- Hirose, O. Acceleration of non-rigid point set registration with downsampling and Gaussian process regression. IEEE Trans. Pattern Anal. Mach. Intell. 43, 2858–2865 (2020).
Google Scholar
- Wang, L. et al. Equivariant local reference frames with optimization for robust non-rigid point cloud correspondence. IEEE Transactions on Image Processing (2025).
- Liu, M. et al. Lcnet: a robust and accurate non-rigid 3d point set registration approach for image-guided liver surgery. IEEE Transactions on Medical Robotics and Bionics (2025).
- Yuille, A. L. & Grzywacz, N. M. A mathematical analysis of the motion coherence theory. Int. J. Computer Vis. 3, 155–175 (1989).
Google Scholar
- Dempster, A. P., Laird, N. M. & Rubin, D. B. Maximum likelihood from incomplete data via the em algorithm. J. R. Stat. Soc.: Ser. B (Methodol.) 39, 1–22 (1977).
Google Scholar
- Tzikas, D. G., Likas, A. C. & Galatsanos, N. P. The variational approximation for Bayesian inference. IEEE Signal Process. Mag. 25, 131–146 (2008).
Google Scholar
- Zeng, H. et al. Integrative in situ mapping of single-cell transcriptional states and tissue histopathology in a mouse model of alzheimer’s disease. Nat. Neurosci. 26, 430–446 (2023).
Google Scholar
- Shi, H. et al. Spatial atlas of the mouse central nervous system at molecular resolution. Nature 622, 552–561 (2023).
Google Scholar
- Zhang, M. et al. Molecularly defined and spatially resolved cell atlas of the whole mouse brain. Nature 624, 343–354 (2023).
Google Scholar
- Wang, Q. et al. The Allen Mouse Brain Common Coordinate Framework: a 3d reference atlas. Cell 181, 936–953 (2020).
Google Scholar
- Schott, M. et al. Open-st: high-resolution spatial transcriptomics in 3d. Cell 187, 3953–3972 (2024).
Google Scholar
- Janesick, A. et al. High resolution mapping of the tumor microenvironment using integrated single-cell, spatial and in situ analysis. Nat. Commun. 14, 8353 (2023).
Google Scholar
- Yoshiura, K. -i et al. A SNP in the ABCC11 gene is the determinant of human earwax type. Nat. Genet. 38, 324–330 (2006).
Google Scholar
- Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn Res 12, 2825–2830 (2011).
Google Scholar
- Wang, L. et al. Extracellular matrix protein 1 (ECM1) is over-expressed in malignant epithelial tumors. Cancer Lett. 200, 57–67 (2003).
Google Scholar
- Rebbeck, C. A. et al. Gene expression signatures of individual ductal carcinoma in situ lesions identify processes and biomarkers associated with progression towards invasive ductal carcinoma. Nat. Commun. 13, 3399 (2022).
Google Scholar
- Cheng, M. et al. Three-dimension transcriptomics maps of whole mouse embryo during organogenesis. bioRxiv https://doi.org/10.1101/2024.08.17.608366 2024–08 (2024).
- Sunkin, S. M. et al. Allen brain atlas: an integrated spatio-temporal portal for exploring the central nervous system. Nucleic Acids Res 41, D996–D1008 (2012).
Google Scholar
- Wang, Y., Liu, Z. & Ma, X. Mnmst: topology of cell networks leverages identification of spatial domains from spatial transcriptomics data. Genome Biol. 25, 133 (2024).
Google Scholar
- Wang, X. et al. Three-dimensional intact-tissue sequencing of single-cell transcriptional states. Science 361, eaat5691 (2018).
Google Scholar
- Zeng, H. et al. Spatially resolved single-cell translatomics at molecular resolution. Science 380, eadd3067 (2023).
Google Scholar
- Wake, H., Lee, P. R. & Fields, R. D. Control of local protein synthesis and initial events in myelination by action potentials. Science 333, 1647–1651 (2011).
Google Scholar
- Zhang, D. et al. Spatial epigenome–transcriptome co-profiling of mammalian tissues. Nature 616, 113–122 (2023).
Google Scholar
- Chen, M., Wei, Z., Huang, Z., Ding, B. & Li, Y. Simple and deep graph convolutional networks. In Proc. of the 37th International Conference on Machine Learning. Vol. 119, 1725–1735 (PMLR, 2020).
- Zhang, H., Wu, Q., Yan, J., Wipf, D. & Yu, P. S. From canonical correlation analysis to self-supervised graph neural networks. Adv. Neural Inf. Process. Syst. 34, 76–89 (2021).
Google Scholar
- Li, L., Li, Z., Zakharchenko, V., Chen, J. & Li, H. Advanced 3d motion prediction for video-based dynamic point cloud compression. IEEE Trans. Image Process. 29, 289–302 (2019).
Google Scholar
- Williams, C. & Seeger, M. Using the nyström method to speed up kernel machines. Adv. Neural Inf. Process. Syst. 13, 661–667 (2000).
- Xiao, Z. et al. 3d reconstruction of a gastrulating human embryo. Cell 187, 2855–2874 (2024).
Google Scholar
- Hu, Y. et al. Benchmarking clustering, alignment, and integration methods for spatial transcriptomics. Genome Biol. 25, 212 (2024).
Google Scholar
- Wolf, F. A., Angerer, P. & Theis, F. J. Scanpy: large-scale single-cell gene expression data analysis. Genome Biol. 19, 1–5 (2018).
Google Scholar
- Wu, T. et al. clusterprofiler 4.0: a universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Google Scholar
- Palla, G. et al. Squidpy: a scalable framework for spatial omics analysis. Nat. Methods 19, 171–178 (2022).
Google Scholar
- Yu, W., Xiaoke, M. & Zaiyi, L. Network model for alignment, stitching and slice-to-volume 3d reconstruction of large-scale spatially resolved slices. https://doi.org/10.5281/zenodo.18815221 (2026).
Download references
Sign in to highlight and annotate this article

Conversation starters
Daily AI Digest
Get the top 5 AI stories delivered to your inbox every morning.
Knowledge Map
Connected Articles — Knowledge Graph
This article is connected to other articles through shared AI topics and tags.

Discussion
Sign in to join the discussion
No comments yet — be the first to share your thoughts!