Assessing uncertainty of sequence representations generated by protein language models
Assessing uncertainty of sequence representations generated by protein language models
Language model-inferred embeddings are replacing structure-derived descriptions of proteins, genes and genomes. We propose a model-agnostic measure to quantify reliability of these new representations.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
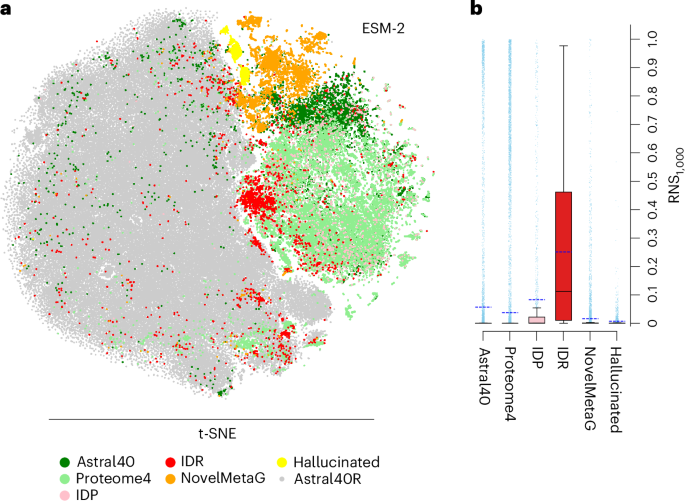
Fig. 1: RNS-based assessments of embeddings identify poorly represented proteins across different data sets.
References
- Vaswani, A. et al. Attention is all you need. Adv. Neural Inf. Process. Syst. 30 (2017). This work introduces the attention mechanism in transformer architecture.
- Weissenow, K. & Rost, B. Are protein language models the new universal key? Curr. Opin. Struct. Biol. 91, 102997 (2025). This review article discusses the transition from evolutionary information to machine-learned embeddings for protein prediction.
Article CAS PubMed
Google Scholar
- Dallago, C. et al. Learned embeddings from deep learning to visualize and predict protein sets. Curr. Protoc. 1, e113 (2021). This article introduces ‘Bioembeddings’, a publicly available library of pLM pipelines.
Article PubMed
Google Scholar
- Saul, B. N. & Christian, D. W. A general method applicable to the search for similarities in the amino acid sequence of two proteins. J. Mol. Biol. 48, 443–453 (1970). The earliest work we have identified that illustrates use of random sequences to evaluate significance of protein sequence similarities.
Article
Google Scholar
Download references
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This is a summary of: Prabakaran, R. & Yana Bromberg, Y. Quantifying uncertainty in protein representations across models and tasks. Nat. Methods https://doi.org/10.1038/s41592-026-03028-7 (2026).
About this article
Cite this article
Assessing uncertainty of sequence representations generated by protein language models. Nat Methods (2026). https://doi.org/10.1038/s41592-026-03027-8
Download citation
- Published: 01 April 2026
- Version of record: 01 April 2026
- DOI: https://doi.org/10.1038/s41592-026-03027-8
Sign in to highlight and annotate this article

Conversation starters
Daily AI Digest
Get the top 5 AI stories delivered to your inbox every morning.
More about
modellanguage model
Speculative decoding works great for Gemma 4 31B in llama.cpp
I get a ~11% speed up with Gemma 3 270B as the draft model. Try it by adding: --no-mmproj -hfd unsloth/gemma-3-270m-it-GGUF:Q8_0 Testing with (on a 3090): ./build/bin/llama-cli -hf unsloth/gemma-4-31B-it-GGUF:Q4_1 --jinja --temp 1.0 --top-p 0.95 --top-k 64 -ngl 1000 -st -f prompt.txt --no-mmproj -hfd unsloth/gemma-3-270m-it-GGUF:Q8_0 Gave me: [ Prompt: 607.3 t/s | Generation: 36.6 t/s ] draft acceptance rate = 0.44015 ( 820 accepted / 1863 generated) vs. [ Prompt: 613.8 t/s | Generation: 32.9 t/s ] submitted by /u/Leopold_Boom [link] [comments]

Gemma 4 - 4B vs Qwen 3.5 - 9B ?
Hello! anyone tried the 4B Gemma 4 model and the Qwen 3.5 9B model and can tell us their feedback? On the benchmark Qwen seems to be doing better, but I would appreciate any personal experience on the matter Thanks! submitted by /u/No-Mud-1902 [link] [comments]

Speed difference on Gemma 4 26B-A4B between Bartowski Q4_K_M and Unsloth Q4_K_XL
I've noticed this on Qwen3.5 35B before as well, there is a noticeable speed difference between Unsloth's Q4_K_XL and Bartowski's Q4_K_M on the same model, but Gemma 4 seems particularly harsh in this regard: Bartowski gets 38 tk/s, Unsloth gets 28 tk/s... everything else is the same, settings wise. This is with the latest Unsloth quant update and latest llama.cpp version. Their size is only ~100 MB apart. Anyone have any idea why this speed difference is there? Btw, on Qwen3.5 35B I noticed that Unsloth's own Q4_K_M was also a bit faster than the Q4_K_XL, but there it was more like 39 vs 42 tk/s. submitted by /u/BelgianDramaLlama86 [link] [comments]
Knowledge Map
Connected Articles — Knowledge Graph
This article is connected to other articles through shared AI topics and tags.
More in Models

Speed difference on Gemma 4 26B-A4B between Bartowski Q4_K_M and Unsloth Q4_K_XL
I've noticed this on Qwen3.5 35B before as well, there is a noticeable speed difference between Unsloth's Q4_K_XL and Bartowski's Q4_K_M on the same model, but Gemma 4 seems particularly harsh in this regard: Bartowski gets 38 tk/s, Unsloth gets 28 tk/s... everything else is the same, settings wise. This is with the latest Unsloth quant update and latest llama.cpp version. Their size is only ~100 MB apart. Anyone have any idea why this speed difference is there? Btw, on Qwen3.5 35B I noticed that Unsloth's own Q4_K_M was also a bit faster than the Q4_K_XL, but there it was more like 39 vs 42 tk/s. submitted by /u/BelgianDramaLlama86 [link] [comments]

Gemma 4 - 4B vs Qwen 3.5 - 9B ?
Hello! anyone tried the 4B Gemma 4 model and the Qwen 3.5 9B model and can tell us their feedback? On the benchmark Qwen seems to be doing better, but I would appreciate any personal experience on the matter Thanks! submitted by /u/No-Mud-1902 [link] [comments]

Kokoro TTS running on-device, CPU-only, 20x realtime!!!
I wanted a reading app where you could read, read and listen or just listen to books with word-by-word highlighting synced to TTS and i wanted the voice to actually sound good. This turned out to be a really hard challenge with Kokoro on iOS, here's what I ran into: Using MLX Swift is great but uses Metal. iOS kills Metal access the moment you background the app. If your use case needs background audio, this is a dead end. ONNX Runtime on CPU fixes the background problem, but the monolithic Kokoro model only runs at 2-3x realtime. After 30 minutes of sustained generation my phone was scorching hot. What actually worked: I split the monolithic model into a multi-stage pipeline and replaced part of the synthesis with native code on Apple's Accelerate framework. That got it to 20x realtime on


Discussion
Sign in to join the discussion
No comments yet — be the first to share your thoughts!